
A new paper, accepted for publication to Journal of Microscopy, has highlighted a 4x increase in acquisition speed whilst also improving image quality in vEM imaging.
Volume imaging techniques in electron microscopy operate by collecting images from sequential serial sections which are then assembled through post-acquisition processing to generate final 3D volume reconstructions. The overall volume that can be included in the final reconstruction is dependent on the uniformity of the sample and the ability of the microscope to image it. This process itself, however, is largely limited by the time to acquire the images and the sample and microscope stability during acquisition. As a result, while vEM is widely used across life sciences and materials research, throughput, reliability and data quality remain key limiting factors.
The paper, published in conjunction with SenseAI, Centre for Ultrastructural Imaging, King’s College London and JEOL, applied workflows from SenseAI to significantly reduce each of these effects and demonstrate significant improvements in acquisition speed, image quality and overall workflow robustness.
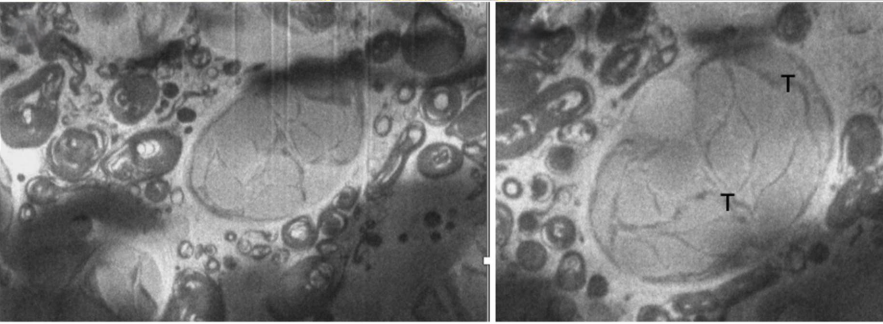
(Left) Paramecium bursaria endosymbiotic alga imaged using SenseAI’s linehop scanning and inpainting algorithm. (Right) Enlargement of symbiotic alga.
The research
Cryo-Focused Ion Beam Scanning Electron Microscopy (cryoFIB-SEM) using samples fixed by high-pressure freezing uniquely enables high resolution cryo-volume Electron Microscope (cvEM) images of cell ultrastructure to be obtained from whole cells and complex tissues in their near native state. cvEM is a leading-edge implementation of volume EM, demonstrating what is possible when acquisition and processing workflows are optimised. As the freezing process also preserves fluorescence, the link between three-dimensional (3D) ultrastructure and biological process is also enabled by targeted cryoCorrelative Light and Electron Microscopy (CLEM). However, the overall viability of advanced vEM workflows such as cvEM is challenged by sample preparation, charge balance during imaging, sample sensitivity to beam damage, contamination, and very long acquisition times. The research tested workflows to significantly reduce each of these effects and demonstrate how both acquisition speed and image quality can be improved simultaneously with results from nematode Caenorhabditis elegans and ciliated protozoon Paramecium bursaria containing many endosymbiotic algae.
Alternative scanning
Each image is typically formed via scanning the electron beam across the face of the sample, in a raster fashion. However, raster scanning can lead to characteristic scanning artefacts due to charging which exist due to the repetitive scanning motion of sweeping the beam from left to right across the sample, whilst always making small movements.
The research employed SenseAI as an alternative scanning method for reducing charging artefacts, and by extension minimising beam damage (without the typical trade-off of reducing dose and degrading signal-to-noise).
SenseAI software works by acquiring only a subset of pixels in a given image, and then applying dictionary learning-based inpainting algorithms to infer the missing data. This approach shifts the imaging paradigm from full acquisition to intelligent sampling and reconstruction.
![]()
Subsampling and inpainting workflow using SenseAI – index and colour indicates order of pixel sampling.
Results
The subsampled scanning from SenseAI demonstrated an increase signal-to-noise ratio for a given dose-budget, whilst also providing smoother contrast and reduction of raster-related artefacts when compared to higher dose examples. There was also a reduction in imaging time that scales linear with the sampling rate – acquiring only 25% of the pixels took only 25% of the time compared to the 100% sampled data. Crucially, reconstructed images retained high-frequency detail and, in many cases, showed improved contrast and interpretability compared to conventional full-dose raster scans. Together, these results demonstrate that faster acquisition does not require compromise: instead, it can deliver both speed and enhanced image quality.
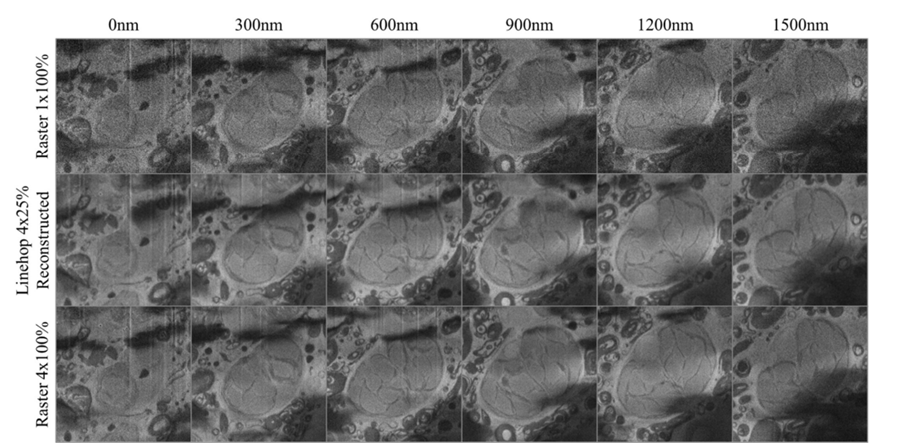

Paramecium bursaria imaged at various cut depths and imaging conditions showing a single endosymbiotic alga. Top row shows a single raster scan acquired with 1us dwell time. Middle row shows a composite image formed from 4 25% linehop scan reconstructions at an equivalent dose level to the top row. Bottom row shows a composite image formed from 4 raster scans. Based on visual clarity, the 4×25% linehop image is very similar to the 4×100% raster scans, indicating a near 4x increase in quality.

